Machine learning & comparative genomics for antimicrobial resistance

Summary
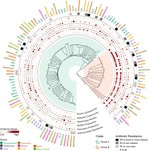
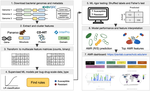
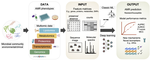
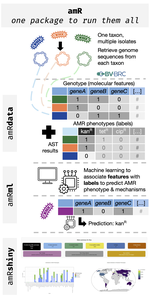
We develop integrative machine learning approaches to predict antimicrobial resistance (AMR) and characterize pathogen genomics across clinical and environmental contexts. Our ML framework associates multi-scale genomic features — genes, protein domains, SNPs, and k-mers — with AMR phenotypes across ESKAPE pathogens, demonstrating that resistance prediction requires integration across sequence scales, with domain-level features often outperforming gene-level markers. Analyses spanning >15,000 genomes reveal that feature importance varies substantially across pathogen species and drug classes, providing mechanistic insights into resistance evolution.
In collaboration with the Penaranda lab (National Jewish Health), we characterized the genetic and phenotypic diversity of Pseudomonas aeruginosa across clinical and environmental isolation sites, revealing distinct genomic signatures associated with niche adaptation and novel resistance determinants. We are extending these approaches to predict host specificity in bacterial pathogens, integrating pangenomic features with host-associated metadata to identify molecular determinants of tissue tropism and species barriers.
Earlier collaborative work with Sreevatsan Lab (while at MSU) identified SNPs in Brazilian Mycobacterium tuberculosis isolates critical for drug resistance, demonstrating how comparative genomics reveals population-specific resistance mechanisms.