MolEvolvR: a web-app for characterizing proteins using molecular evolution and phylogeny
Abstract
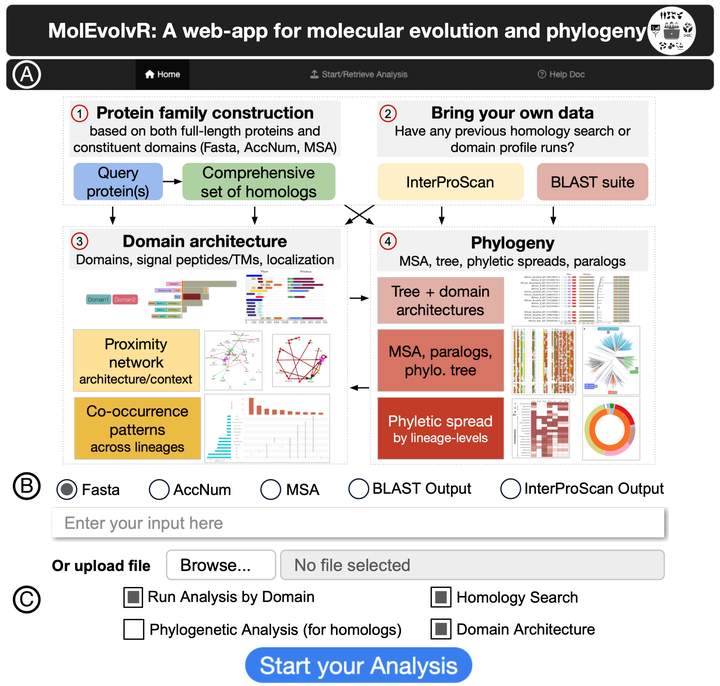
Studying proteins through the lens of evolution can reveal features such as conserved domains, lineage-specific variants, and co-occurring domain architectures in phylogenetic context across all superkingdoms. MolEvolvR enables researchers to conduct such evolution-focused studies to generate testable hypotheses about protein function and evolution. MolEvolvR is a novel web-app allowing researchers to visualize the molecular evolution of their proteins of interest in a phylogenetic context across the tree of life. It accepts multiple input formats – protein/domain sequences, homologous proteins, or domain scans – and, using a general-purpose computational workflow, returns detailed homolog data and dynamic graphical summaries (e.g., phylogenetic trees, multiple sequence alignments, domain architectures, domain proximity networks, phyletic spreads, co-occurrence patterns across lineages). MolEvolvR performs domain-centric searches to capture remote homologs that are missed by full-length searches, integrates domain architecture evolution with phyletic distribution analyses, and provides evolutionary context visualizations that reveal lineage-specific adaptations versus those that are broadly conserved. Thus, MolEvolvR is a powerful, easy-to-use web interface for computational protein characterization. The web-app can be accessed here: https://jravilab.org/molevolvr.
FSA, JTB, LS are co-primary authors who contributed equally; they are listed alphabetically.