Dynamical systems & modeling
Summary
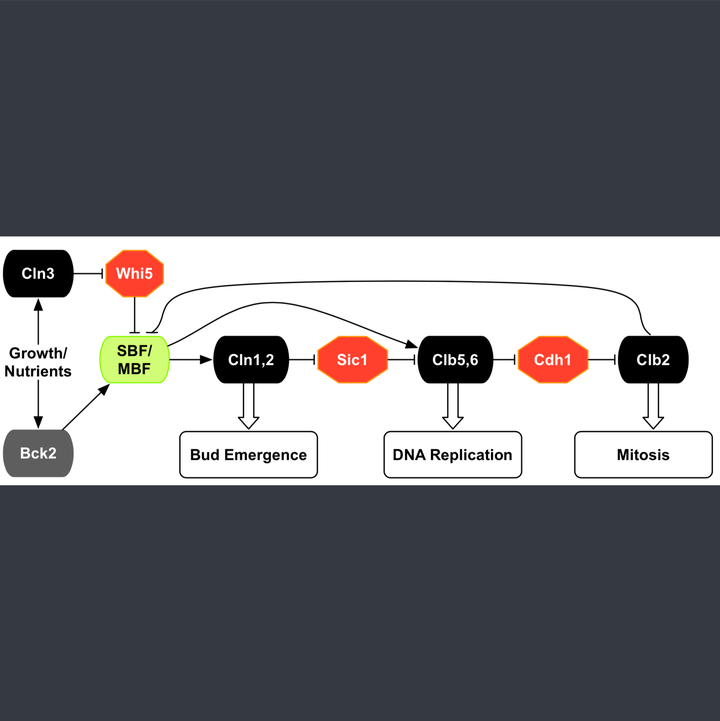
We developed mathematical models that capture dynamic aspects of cellular regulatory systems. A detailed model (~100 molecular species, ~150 parameters) of the START transition in budding yeast reconciles diverse experimental findings on mechanisms and timing of transcriptional, post-translational, and localization events, and addresses the nutritional effect on size control. Consistent with ~200 experimental phenotypes, the model successfully predicts whether novel mutants rescue or induce cell cycle arrest; many predictions have been experimentally validated. Interactive model simulations are available for the research community to explore START dynamics.
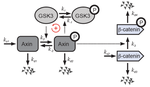
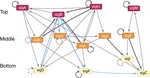
We also built a simplified model of the Wnt signaling pathway demonstrating that incorporating key feedback regulatory interactions enables bistable behavior, reconciling theoretical predictions with experimental observations of cellular memory in differentiation decisions. In bacterial systems, network analysis of the M. tuberculosis sigma factor regulatory network revealed a master regulator-initiated three-tiered hierarchy and distinct sigma factor communities with over-represented network motifs including autoregulation and coregulation of sigma/anti-sigma pairs.
+++