Lab News
New NIH R21 grant on phage-bacteria interactions!
2025–2026 Manuscripts
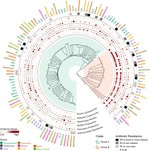
- Penaranda C^, Brenner EP^, Clatworthy AE, Cosimi LA, Ravi J*, Hung DT*. Genomic comparison and phenotypic characterization of Pseudomonas aeruginosa isolates across environmental and diverse clinical isolation sites. mSystems. 2026. DOI. Editor’s pick.
- [preprint] Brenner EP^, Vang CK^, Johnson CG, Ravi J*. Genotype-phenotype modeling of light ecotypes in Prochlorococcus reveals genomic signatures of ecotypic divergence. bioRxiv. 2025. DOI
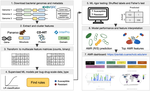
- [preprint] Ghosh A^, Brenner EP^, Vang CK^, Wolfe EP^, Boyer EA, Lesiyon RL, Manpearl KR, Sridhar V, Burke JT, Krol JD, Bilodeaux JMR, Ravi J*. From sequence to signature: Machine learning uncovers multiscale feature landscapes that predict AMR across ESKAPE pathogens. bioRxiv. 2025. DOI
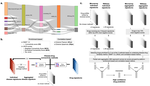
- [preprint] Samart K, Thang L, Buskirk LR, Tonielli AP, Krishnan A*, Ravi J*. Integrative transcriptome-based drug repurposing in tuberculosis. bioRxiv. 2025. DOI
- Ghosh A^, Vang CK^, Brenner EP^, Ravi J*. Unlocking antimicrobial resistance with multiomics and machine learning. Trends in Microbiology. 2025.
DOI
^co-primary authors; *co-corresponding authors. In bold, members of JRaviLab.
NIH Bridge2AI U54 & DOE BRaVE grants!
We are co-investigators on two new interdisciplinary grants:
- (co-I) NIH Common Fund, Bridge to Artificial Intelligence (Bridge2AI) U54 | Integration, dissemination, and evaluation center for the NIH Bridge2AI program. 2025–2026.
- (co-I) Department of Energy, Biopreparedness Research Virtual Environment (BRaVE) Award to PNNL | Enhancing biopreparedness through a model system to understand the molecular mechanisms that lead to pathogenesis and disease transmission. 2024–2026.
2024 Manuscripts
- Janani Ravi, Vivek Anantharaman, Samuel Z Chen, Evan P Brenner, Pratik Datta, L Aravind, Maria L Gennaro. The phage shock protein (PSP) envelope stress response: discovery of novel partners and evolutionary history. mSystems. 2024. DOI.
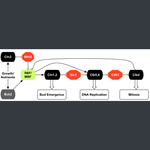
- Janani Ravi, Kewalin Samart, Jason Zwolak. Modeling the START transition in the budding yeast cell cycle. PLoS Comp Biol. 2024. DOI. *Selected for cover art of the Aug 2024 issue.
Translational Research Scholars Program!
CPMR-DBMI dyad pilot grants!
We received two Colorado Program for Musculoskeletal Research (CPMR) / Department of Biomedical Informatics (DBMI) dyad pilot grants.
- With Dr. Honey Hendesi on studying how the effect of intermittent fasting on the gut microbiome impacts bone health and fracture healing
- With Dr. Reed Ayers on studying metallosis — how bacteria (and hosts) break down metal implants (that supposedly can’t be corroded!)
See Associated news article
NIH U01 funding!
Our project on developing novel data science technologies to study antimicrobial resistance is officially now funded by NIH NIAID! If you are interested in bridging microbial genotypes to phenotypes, esp. in the context of AMR, come join us!
Now looking for postdocs: check out CU ad here.
New folks in the group! (2022-2023)
Exciting times as we welcome new people to our group:
- 2023-06: Evalyn Resnick, undergraduate researcher (planning to attend Simpson College starting in the Fall of 2023; Biology major – pre-medicine/pre-veterinary track)
- 2023-06: Kritika Verma, undergraduate researcher (pursuing Bachelor of Technology, Cluster Innovation Centre at University of Delhi)
- 2023-06: Sana Paktinyar, undergraduate researcher (pursuing BS in Bimolecular Chemical Engineering, Pre-Med Track at California Institute of Technology)
- 2023-06: Vy ‘Andy’ Tran, undergraduate researcher (pursuing BS in Physics at University of Colorado Boulder)
- 2022-11: Jake Krol, postbac, Informational Science Professional (BS, Computational Neuroscience from MSU)
- 2023-02: Karla Vela Lopez, graduate researcher (pursuing MS in Biomedical Sciences and Biotechnology, CU Anschutz)
- 2023-02: Jill Bilodeaux, graduate rotation student from Microbiology
- 2023-03: Kewalin Samart & Keenan Manpearl, two graduate pseudo-rotation students from Computational Biosciences
- 2023-05: Evan Brenner, postdoc (PhD, Comparative Medicine and Integrative Biology, MSU)
2023 Manuscripts
- Joshua M. Lensmire, Michael R. Wischer, Cristina Kraemer-Zimpel, Paige J. Kies, Lo Sosinski, Elliot Ensink, Jack P. Dodson, John C. Shook, Phillip C. Delekta, Christopher C. Cooper, Daniel H. Havlichek Jr., Martha H. Mulks,Sophia Y. Lunt, Janani Ravi, Neal D. Hammer. The glutathione import system satisfies the Staphylococcus aureus nutrient sulfur requirement and promotes interspecies competition. PLoS Genetics . 2023. DOI.
Highlighted in CU Anschutz newsletter
Moved to CU Anschutz
 We are thrilled to announce that the JRaviLab recently moved from MSU to CU Anschutz. We are now part of the new Dept. of Biomedical Informatics, Center for Health Artificial Intelligence, University of Colorado Anschutz.
We are thrilled to announce that the JRaviLab recently moved from MSU to CU Anschutz. We are now part of the new Dept. of Biomedical Informatics, Center for Health Artificial Intelligence, University of Colorado Anschutz.First NIH $$
- (co-PI) Notice of Award for NIH NIAID Data Science R21 on “Mechanism-guided drug repurposing for host-directed therapy of infectious diseases using interpretable and integrative ML” | co-PI: Arjun Krishnan
2022 Student Accomplishments
- Ethan receives a summer research scholarship from the MSU College of Natural Science, and presents at his first international meeting!
- [Outgoing] Kewalin graduated from MSU with the McCartney Math Award for outstanding graduating Math majors. She will be joining the University of Colorado, Anschutz Medical Campus, Computational Biosciences graduate program this Fall (joining us back in Aug @CU!). She also received grad admits at MSU (BMS, CMSE), and Penn State, along with the College of Engineering Distinguished Scholarship at MSU, the Rasmussen Doctoral Recruitment Award at MSU.
- [Outgoing] Joseph Burke graduated from MSU with an outstanding student award from the Dept of Microbiology and Molecular Genetics, and three co-primary author publications under his belt. He’s now a software engineer at CRISPR Therapeutics.
- [Outgoing] Elliot Majlessi graduated from MSU and is taking a gap year to apply to Medical School!
- [Outgoing] Mary Zhang graduated from Carleton College to join med school at the National University of Singapore, Fall of ‘22!
2022 Manuscripts
- Burke JTp, Chen SZp, Sosinski LMp, Johnston JB, Ravi Jc. MolEvolvR: A webapp for characterizing proteins using molecular evolution and phylogeny. bioRxiv 2022. Webapp
- Joo Rp,c, Sánchez-Tapia Ap, Mortara S, Saibene YB, et al., Ravi Jc. Ten simple rules to host an inclusive conference. PLoS Computational Biology. 2022. DOI GitHub
- Conner KNp, Burke JTp, Ravi Jc, Hardy JWc. Novel Internalin P homologs in Listeria ivanovii londoniensis and Listeria seeligeri. Microbial Genomics. 2022. DOI GitHub
- Blaine HCp, Burke JTp, Ravi Jc, Stallings CLc. DciA helicase operators exhibit diversity across bacterial phyla. Journal of Bacteriology. 2022. DOI GitHub
- Cantoria MJp, Alizadeh Ep, Ravi J, et al., Feedback in the β-catenin destruction complex imparts bistability and cellular memory. bioRxiv 2022.
- [update] Hsueh BYp, Severin GBp, et al., now at Nature Microbiology (2022)!
DOI
pco-primary authors; cco-corresponding authors. In bold, members of JRaviLab.
2021 Manuscripts
- [update] Samart Kp, Tuyishime Pp, Krishnan Ac, Ravi Jc published in Briefings in Bioinformatics (2021)! DOI | Live doc | GitHub.
- Ravi Jp,c, Fioravanti Ap,c. S-layers: the proteinaceous multifunctional armours of Gram-positive pathogens. Frontiers in Microbiology 2021. DOI
- Hsueh BYp, Severin GBp, et al., A Broadly Conserved Deoxycytidine Deaminase Protects Bacteria from Phage Infection. bioRxiv 2021.
- Lensmire JM, et al., Ravi J, Hammer ND. The glutathione import system satisfies the Staphylococcus aureus nutrient sulfur requirement and promotes interspecies competition. bioRxiv 2021.
- [update] Hadi SAp, Kolte IVp, Brenner EPp, et al., Ravi J, Sreevatsan S, Basta PC. Identification of a predominant genotype of Mycobacterium tuberculosis in Brazilian indigenous population. Scientific Reports. 2021;11(1):1224.
DOI
pco-primary authors; cco-corresponding authors. In bold, members of JRaviLab.
More $$ for the group!
- (PI) Awarded the Endowed Research, Sheila McMonagle Fund from the College of Veterinary Medicine, MSU for the molecular evolution project!
- (PI) Received grant support from the NSF-funded BEACON Center for ML-based feature detection and comparative pathogenomics.
- (co-I) Received a grant from Spectrum Health and MSU Alliance for a COVID-19 project!
- (subaward) Received NIH NIAID grant subaward to study molecular pathogenesis in Staphylococci using computational approaches
2021 Student Accomplishments
- Kewalin, Sam, Vignesh, Elliot, and Joe present at their first (inter)national meetings!
- Kewalin received several research scholarships for 2021, including the Engineering Summer Undergraduate Research Experience (EnSURE), MSU; College of Natural Science scholarships for Spring and Fall.
- Ethan Wolfe and Amy Tonielli join the group! They received Professorial Assistantships from the Honors College, MSU (given to the Top200 students).
- Elliot received the NIH R25-funded BRUSH summer scholarship.
- [Outgoing] Sam Chen graduated as one of 38 students across MSU with a 4.0 GPA. He’s now a software engineer at Amazon Inc.
- [Outgoing] Phil Calhoun graduated with Honors and joined the MSU College of Human Medicine.
First manuscripts submitted from JRaviLab!
Two preprints submitted to arXiv and bioRxiv in Sep ‘20! Another manuscript under review in Scientific Reports w/ collaborator, Dr. Sreevatsan!
- Samart Kp, Tuyishime Pp, Krishnan Ac, Ravi Jc. Reconciling multiple connectivity scores for drug repurposing. arXiv 2020 | Live doc | GitHub.
- Ravi Jc, Anantharaman V, Chen SZ, Datta, P, Aravind Lc, Gennaro MLc. Phage-shock-protein (Psp) Envelope Stress Response: Evolutionary History & Discovery of Novel Players.
bioRxiv 2020 |
Webapp.
pco-primary authors; cco-corresponding authors. In bold, members of JRaviLab.
2020 Student Accomplishments
- Karn and Sam received Honorable mention for their posters at ISMB 2020 (by International Society for Computational Biology)!
- Karn and Sam were nominated for the NSF-funded REU-ACRES program from ICER, MSU! Karn also gets accepted into the AdvanceU internship program at MSU!
- Phil received the NIH NHLBI-funded BRUSH scholarship from the College of Veterinary Medicine!
- Kewalin received the MSU College of Natural Science, Scholarship to work with us this Spring!
- Lauryn joined us from Hampton University, Virginia starting Jul ‘20 (as part of NSURP, BIPOC research program for microbiologists). Now a PREP student at the University of Pennsylvania.
- [Outgoing] Phoebe received the Mastercard foundation scholarship to work back home in Rwanda from Oct ‘20; University of British Columbia, Fall ‘21. She also gets grad admits from Purdue University and UC Davis!
- [Outgoing] Lo got accepted into the MSU post-baccalaureate program to start working with Dr. Rob Quinn (BMB, MSU) from Sep ‘20. Lo is now pursuing their PhD at Michigan State University.
Janani & Camille win the Excellence in Diversity Award!
2019 Grants
 Received the Launch Award from MSU’s Diversity Research Network for the drug-repurposing project!
Received the Launch Award from MSU’s Diversity Research Network for the drug-repurposing project! Awarded the Endowed Research, Harvey J Fiege Fund from the College of Veterinary Medicine, MSU for the molecular evolution project!
Awarded the Endowed Research, Harvey J Fiege Fund from the College of Veterinary Medicine, MSU for the molecular evolution project!