Bacterial stress response
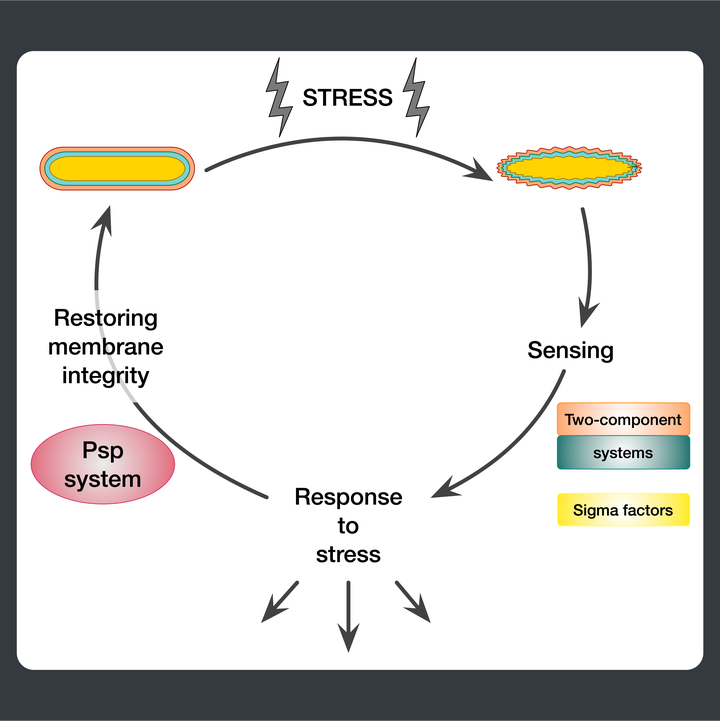
Summary
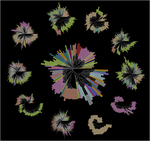
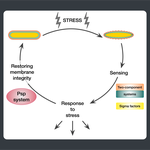
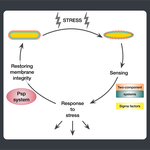
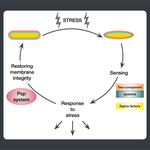
Starting from postdoctoral research at Rutgers University, we have used computational evolutionary approaches to study the phage shock protein (PSP) envelope stress response system across the tree of life. Our work revealed that the minimal PSP system was present in the last universal common ancestor — with its core protein PspA (Snf7 in ESCRT outside bacteria) diversifying into multiple distinct PSP systems across life. Systematic mapping of domain architectures and genomic contexts uncovered novel partners including the Toastrack domain, HTH-associated signaling domain, and ATPase and Band-7 domain associations.
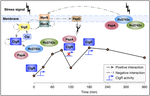
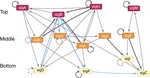
Earlier work delineated the variations of the Psp system across Actinobacteria, and reconstructed and topologically characterized the sigma factor regulatory network of M. tuberculosis, revealing a master regulator-initiated three-tiered hierarchy with over-represented network motifs including autoregulation and coregulation of sigma/anti-sigma pairs. Results are available at our interactive PSP web app.