Molecular evolution & comparative pathogenomics
Summary
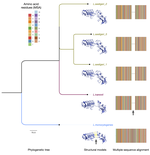
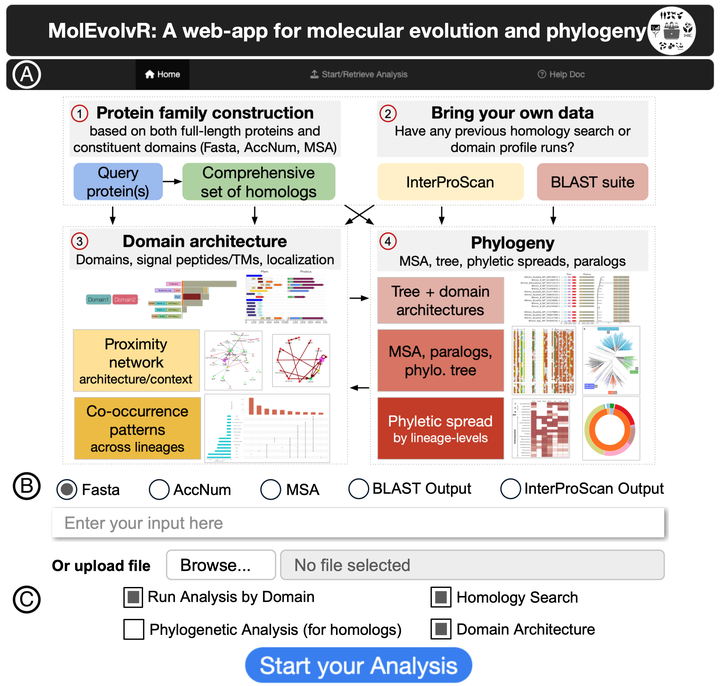
We developed MolEvolvR, a web application that integrates sequence analysis, domain architecture mapping, phylogenetic reconstruction, and genomic context visualization to enable systematic study of protein family evolution across the tree of life. MolEvolvR has engaged 45+ trainees worldwide through the Outreachy open-source program; we are developing MolEvolvR 2.0 with a companion R package for programmatic access.
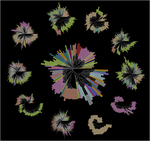
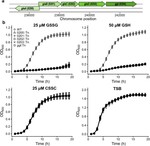
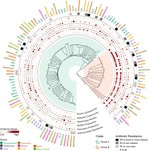
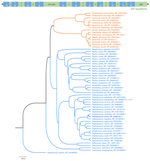
Our foundational work on the phage shock protein (PSP) envelope stress response exemplifies this approach: discovering that a minimal PSP system was present in the last universal common ancestor and tracing its diversification into novel functional contexts across life, including novel partners (Toastrack domain, HTH-associated signaling domain, ATPase and Band-7 domain associations). We have applied this evolutionary framework broadly to characterize protein families across diverse pathogens — including bacterial phage defense via deoxycytidine deaminases, cell surface S-layer proteins in Gram-positive pathogens, internalins in Listeria, DciA helicase loaders across bacterial phyla, and sulfur acquisition systems in Staphylococcus aureus. This work is foundational to our group’s philosophy of using evolutionary genomics to predict protein function with both basic biology and translational impact.